Antibodies against PTM
PTM specific antibodies applications
Know more about the usage and issues of PTM-antibodies to evaluate specific enzymatic activities, with a strong focus on kinases and methylases.
Use of PTM-specific antibodies
in end users’ applications
Kinases, methylases and acetylases
The proteins are modified post-translationally by enzymes, such as kinases, methylases and acetylases. It is well known that such enzymes are not specific to one substrate only; for example, kinases such as MAP kinases, JAK, PI3K and many others, when activated upon a specific stimulus in a specific cellular context, will not phosphorylate one target only, but usually a subset of different protein targets which will collectively lead to the expected stimulation.
Kinases consensus sequences
Each kinase will phosphorylate its substrates on amino acid patterns sharing sequence similarities, and many publications describe the consensus sequences recognized by particular kinases. Therefore, the sequence surrounding one PTM site in one protein will share sequence features with other proteins, all being the substrates of a same kinase (‘kinase substrate family’). Therefore, when raising an anti-PTM antibody to one protein from a family, the risk is high that this antibody will also cross-react to other family members in the same sample. This cross-reactivity is somehow expected, therefore, and cannot be considered as a non-specific reaction.
Anti-PTM antibodies
To determine if the anti-PTM antibody does recognize the protein member in a specific assay, it is advised to perform a western blotting experiment, and refer to the size of the target protein member to assess its phosphorylation state.
Success story
Deciphering the histone code
According to the ‘histone code’ hypothesis, combinatorial modifications dictate a biological readout.
Phosphorylation of serine residues within Histone H3 is a highly dynamic process that associates with acetylation and methylation to create combinatorial patterns which are read by specific detector proteins (A. Sawicka & C. Seiser (2012) Biochimie 94, 2193-2201). a model has been proposed, where 14-3-3 binding to H3S10ph at target promoters is stabilized by additional acetylation of H3K9 or H3K14.
Dr. Christian Seiser (Medical University of Vienna) used the Speedy protocol to develop a modification specific antibody.
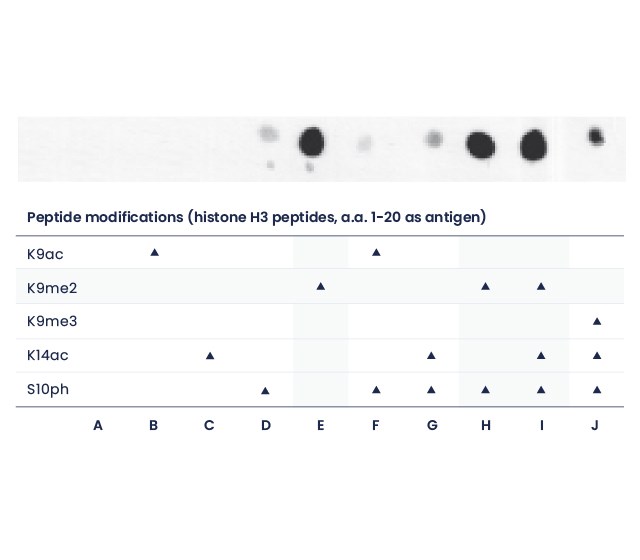
Dot blot
A polyclonal antibody targeting the dimethyl-Lysine 9 of Histone 3 was tested against various H3 peptides and shown to bind to all peptides containing the dimethyl-Lysine 9, no matter the surrounding residues.
Dot blot (1 µg peptides spots)
The Speedy 28-day polyclonal antibody recognizes Histone H3 K9me2 (E), independently of neighboring modifications S10ph and K14ac marks (H,I). The antibody specificity has been determined with different peptides derived from histone H3 (amino acids 1-20).
In situ Hybridization assay
The polyclonal anti-H3 K9me2 is able to recognize Histone H3 multi-modified on K9me2 S10ph K14ac.
In situ Hybridization assay
Swiss 3T3 fibroblasts were stimulated with anisomycin/TSA (left), or left untreated (right). The cells were fixed, blocked, and incubated with the purified polyclonal antibody to K9me2. The detection was done with Alexa 595®-conjugated secondary anti-rabbit antibodies.