COVID-19
Inside the SARS-CoV-2 Genome
SARS-CoV-2 genome sequence, phylogeny data and virus evolution are crucial information to understand how the virus operates. An accurate knowledge of the virus genome guarantees the development of efficient and specific diagnostic tests and preventive and curative treatments.
SARS-CoV-2 genome
SARS-CoV-2 genome is a positive-sense single-stranded RNA of 29.9 kb.
Genome sequencing indicates that the virus genome code for 10 genes generating 29 proteins.
Genome organization
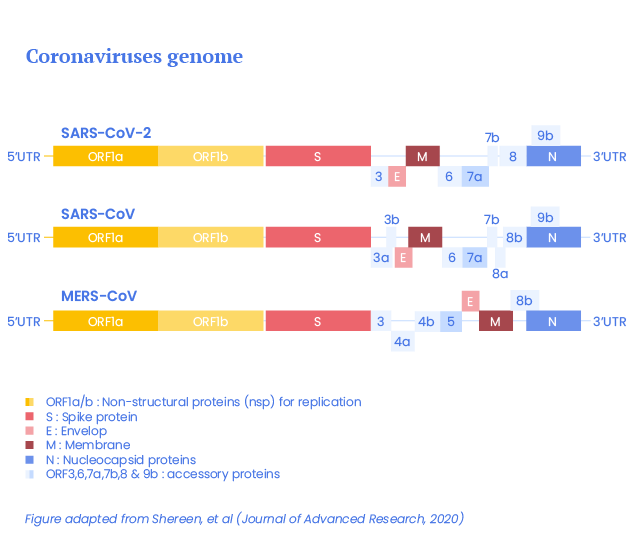
The virus genome is constituted of the following genes :
- a 5’ UTR (untranslated region) cap,
- replicase complex of ~20 kb (ORF1a and ORF1b),
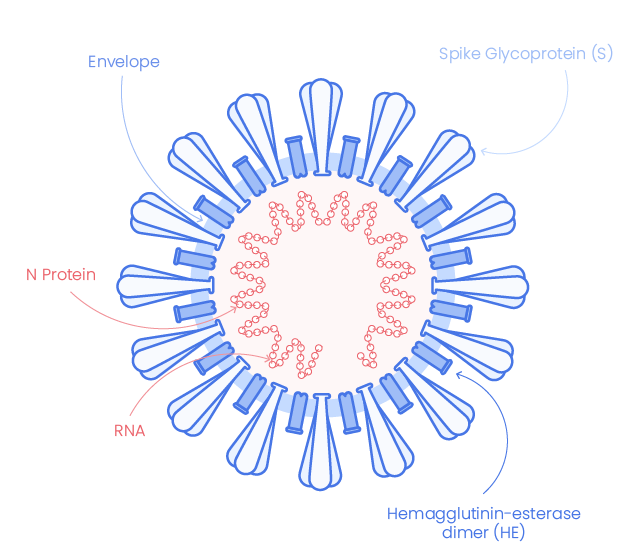
- S gene (coding for the spike protein – 1273 aa),
- E gene (coding for the envelop protein – 75 aa),
- M gene (coding for the membrane protein – 222 aa),
- N gene (coding for the nucleocapsid proteins – 419 aa),
- 3’ UTR including a poly-A sequence and
- unidentified open reading frames (ORF). (1)
ORF1a is translated directly from the genomic RNA into 11 non-structural proteins while ORF1b expression requires a ribosomal frameshift to generate 5 non-structural proteins. (2)
SARS-CoV-2 Life Cycle
Learn more about the coronavirus life cycle and discover products
to help you in your researches.
Genome evolution
of 2019-nCoV
It has been reported that 2 genotypes of SARS-CoV-2 do exist. The S and L types named according to the amino acid 84 (S84L). S and L types can be differentiated by 2 SNPs at positions 8782 (orf1ab: T8517C) and 28144 (ORF8: C251T, S84L). S genotype SARS-CoV-2 is described to be the ancestral version while the L-type evolved from the S-type and seems to have a higher frequency. It is still unclear why L-type is more virulent and how it evolved from the S-type. (4)
A SARS-CoV-2 variant with a 382 nucleotides deletion emerged in Wuhan early in the pandemic. The deletion occurred in ORF7b removing the ORF8 transcription regulatory sequence.
This led to virus attenuation and reduced replication but lead to immune evasion. ORF8 is considered a hotspot for genetic variation. Other variants with a deletion in ORF8 were reported in Bangladesh (345 nucleotides), Australia (138 nucleotides), and Spain (62 nucleotides) (5)
Studies are continuously made to keep identifying the emergence of new variants of SARS-CoV-2. Sequence analysis is a key point to ensure testing efficacy if a change occurs in immune targets or oligonucleotide (primers, probes) binding sites.
Decipher the SARS CoV codes
Decoding the SARS-CoV-2 genome implies sample collection and analysis.
Viral RNA analysis and phylogenetic studies call for genomic related reagents
such as oligonucleotides, polymerases, and electrophoresis related products.
Choose your weapons
We are experts in the production of oligonucleotides and have a large portfolio of usual and exotic modifications. We offer a large panel of fluorophores and quenchers covering all the qPCR channels. Our Access™ Dyes and Quenchers range offers a class of proprietary molecules or other dyes & quenchers with unmatched detection performance and IP-friendly terms and conditions.
For sequencing purposes, we recommend our NGS custom oligonucleotides specially produced to avoid cross-contamination and reduce barcode misalignment during multiplex next-generation sequencing projects.
| RUO Oligonucleotides | Configure |
|---|---|
| Diagnostic Oligonucleotides | Request a quote |
| Therapeutic Oligonucleotides | Request a quote |
| NGS Oligonucleotides | Configure |
| Double-Dye Probes | Configure |
| Molecular Beacons | Configure |
| MGB Probes | Configure |
| Access™ Dyes and Quenchers | View our Offer |
Small interference RNAs (siRNAs) are potential candidates for anti-SARS drugs. siRNA could be used to target key genes and block virus replication. We provide ready-to-use siRNA Controls and custom synthesis of siRNA Duplexes. We can assist you with the design of your sequences.
For customers who prefer to synthesize by themselves their nucleic acid sequences, we distribute synthesis reagents from Glen Research company. Glen Research is a provider of critical reagents required during the COVID pandemic. As a distributor of their products in most European countries, we can help you in synthesizing high-quality oligonucleotides.
SARS-CoV-2 related subjects
References
1) A novel coronavirus from patients with pneumonia in China, 2019.
Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, et al.
N Engl J Med. 2020;382(8):727–33 – DOI: 10.156/NEJMoa2001017
2) Genome evolution of SARS-CoV-2 and its virological characteristics.
So Nakagawa and Takayuki Miyazawa.
Inflammation and Regeneration (2020) 40:17 – DOI: 1186/s41232-020-00126-7
3) A pneumonia outbreak associated with a new coronavirus of probable bat origin.
P. Zhou, X.L. Yang, X.G. Wang, B. Hu, L. Zhang, W. Zhang, H.R. Si, Y. Zhu, B. Li, C.L. Huang, H.D. Chen, J. Chen, Y. Luo, H. Guo, R.D. Jiang, M.Q. Liu, Y. Chen, X.R. Shen, X. Wang, X.S. Zheng, K. Zhao, Q.J. Chen, F. Deng, L.L. Liu, B. Yan, F.X. Zhan, Y.Y. Wang, G.F. Xiao, Z.L. Shi.
Nature 579 (7798) (2020) 270–273 – DOI: 1038/s41586-020-2012-7
4) On the origin and continuing evolution of SARS-CoV-2.
Tang X, Wu C, Li X, Song Y, Yao X, Wu X, et al.
Natl Sci Rev. 2020 Mar 3 : nwaa036 – DOI: 1093/nsr/nwaa036
5) Effects of a major deletion in the SARS-CoV-2 genome on the severity of infection and the inflammatory response: an observational cohort study.
Barnaby E Young et al.
Vol 396, August 18, 2020 - DOI: 10.1016/ S0140-6736(20)31757-8